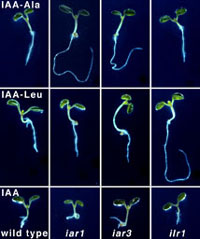
Isolation and characterization of Arabidopsis mutants allowed us to identify genes providing inputs to the auxin pool, to distinguish precursors from active auxin, and to disentangle contributions of different auxin biosynthetic pathways to development. We isolated a collection of Arabidopsis mutants resistant to auxin
storage forms (including the IAA-amino acid conjugates IAA-Ala
and IAA-Leu) that remained sensitive to free IAA.
Two of these mutants, ilr1 and iar3, led to the discovery of enzymes that can release free
IAA from conjugates. In contrast, ilr2, ilr3,
iar1, and mtp5 defects uncovered a role for metal homeostasis
in auxin metabolism.
Our auxin conjugate publications
|
Lab members with conjugate projects:
Collaborator:
Rebekah Rampey
Former Graduate Students:
Haifeng Chen (M.A., 1999)
Jamie Lasswell (Ph.D., 2000)
Sherry LeClere (Ph.D., 2002)
Mónica Magidin (Ph.D., 2002)
Rebekah Rampey (Ph.D., 2004)
Andrew Woodward (Ph.D., 2005)
Luise Rogg (Ph.D., 2001)
Rosie Tellez (1995-2001)
We gratefully acknowledge support
for this research from the NIH, the Robert A. Welch Foundation,
a NIH Training Grant (T32-GM08362; SL), and Houston Livestock
Show and Rodeo Scholarships (RR, SL, JL, RT).

|